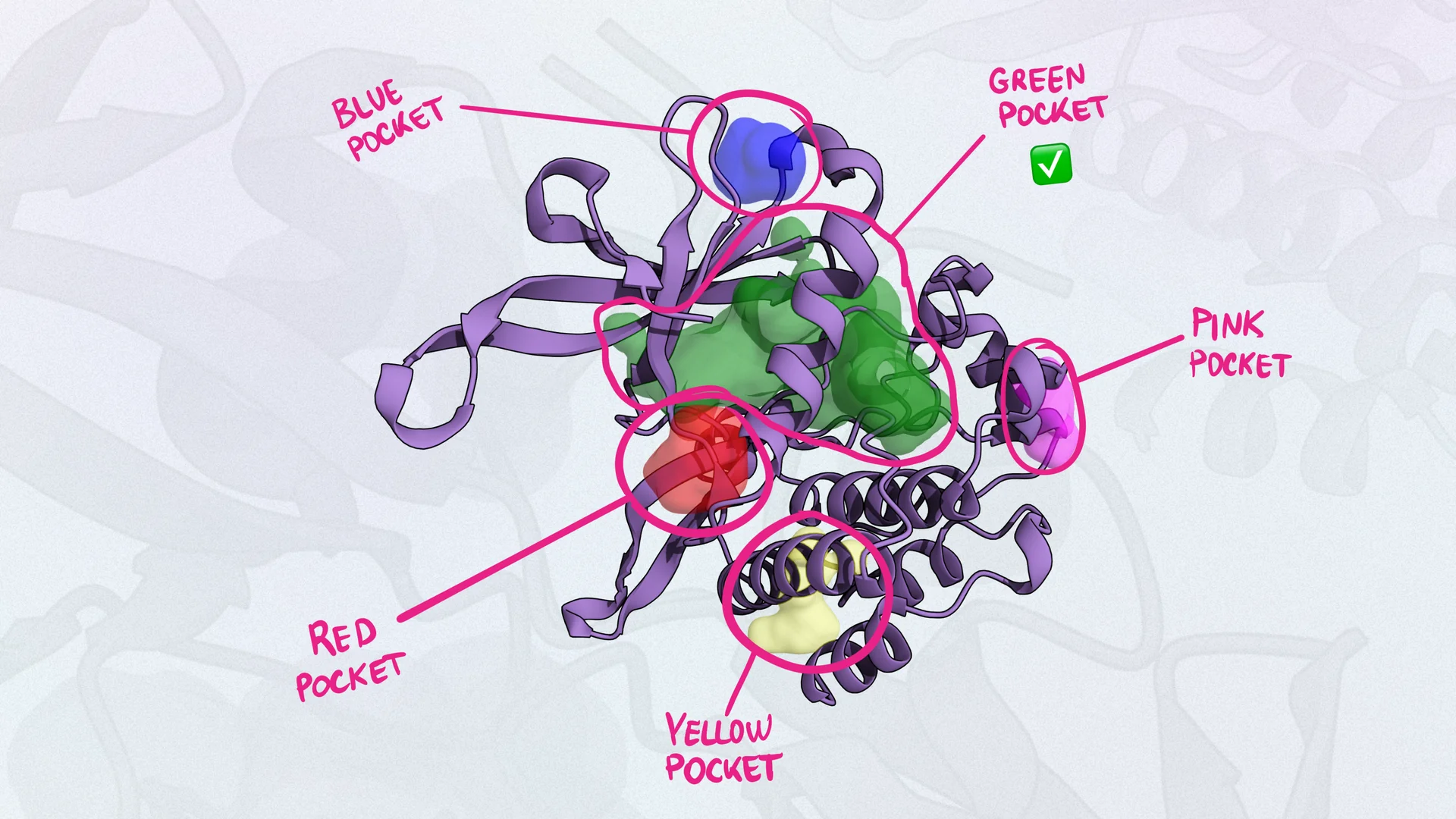
Docking
methodMolecular docking is a computational technique used to predict the preferred orientation of a ligand to a target (usually to a pre-defined site). This process helps to predict the strength and nature of the interactions between the molecules. It typically involves two steps: I) sampling different poses, II) assessing the pose through a scoring function that describes the energetic contribution of protein-ligand interactions.
Importance in Computational Drug Discovery
- Binding Pose Prediction: Docking can predict how and where a ligand binds to a protein, which is crucial for understanding the molecular basis of the interaction.
- Binding Affinity Estimation: Docking scores provide an estimate of the binding affinity, helping to identify potential ligands with strong interactions.
- Structure-Based Drug Design: Docking facilitates the design of new molecules by modeling how structural changes in the ligand or target affect binding.
- Virtual Screening: High-throughput docking allows the screening of large libraries of compounds to identify potential ligands.
Key Tools
DeepOrigin Dock: A tool predicting binding affinities and docking poses using advanced algorithms and integration with other drug discovery tools. Available via Balto and API.
Dock 3.8: A computational tool that performs structure-based virtual screening by docking small molecules into protein binding sites to predict binding poses and affinities.
AutoDock Vina: An open-source program for molecular docking and virtual screening.
AlphaFold 3: A tool for predicting protein structures and protein-ligand complexes enabling structure-based coking and facilitation of modeling of molecular interactions.
Literature
-
“Molecular Docking in Drug Discovery”
-
Publication Date: 2021-06-03
-
Summary: Discusses molecular docking methods for optimizing lead molecules, evaluating biological pathways, de novo drug design, and structure-based virtual screening.
-
“Molecular Docking in Drug Discovery: Techniques, Applications, and Advancements”
-
Publication Date: 2024-10-16
-
Summary: Reviews various approaches and methods used in molecular docking, and techniques for interpreting and validating docking results.
-
“Molecular Docking: Shifting Paradigms in Drug Discovery”
-
Publication Date: 2019-09-01
-
DOI: 10.3390/ijms20184331
-
Summary: Describes initial applications of molecular docking and newer uses, including prediction of adverse effects, polypharmacology, drug repurposing, and target profiling.
-
“Molecular Docking: Principles, Advances, and its Applications in Drug Discovery”
-
Publication Date: 2022-09-22
-
Summary: Reviews the principles and advances in molecular docking and its applications in drug discovery.
-
“Review on the use of Molecular Docking as the First Line Tool in Drug Discovery and Development”
-
Summary: Discusses different molecular docking methods, software used, and their applications in drug discovery.
-
“Ensemble Docking in Drug Discovery: How Many Protein Configurations from Molecular Dynamics Simulations are Needed To Reproduce Known Ligand Binding?”
-
Publication Date: 2019-01-29
-
Summary: Demonstrates the use of molecular dynamics simulations to generate protein configurations for docking campaigns.
-
“Applications of Molecular Docking in Natural Products-Based Drug Discovery”
-
Publication Date: 2023-02-01
-
Summary: Discusses molecular docking in natural product drug discovery programs.
-
“Virtual Screening, Molecular Docking and QSAR Studies in Drug Discovery and Development Programme”
-
Publication Date: 2020-07-15
-
Summary: Comprehensive review on computational tools such as virtual screening, molecular docking, and QSAR methods in drug discovery.
-
“Theory and Applications of Covalent Docking in Drug Discovery: Merits and Pitfalls”
-
Publication Date: 2015-01-27
-
Summary: Highlights different aspects of covalent docking, its merits, and pitfalls.
-
“Consensus Docking in Drug Discovery”
-
Publication Date: 2020-03-31
-
Summary: Discusses consensus docking approaches to improve docking and virtual screening results.