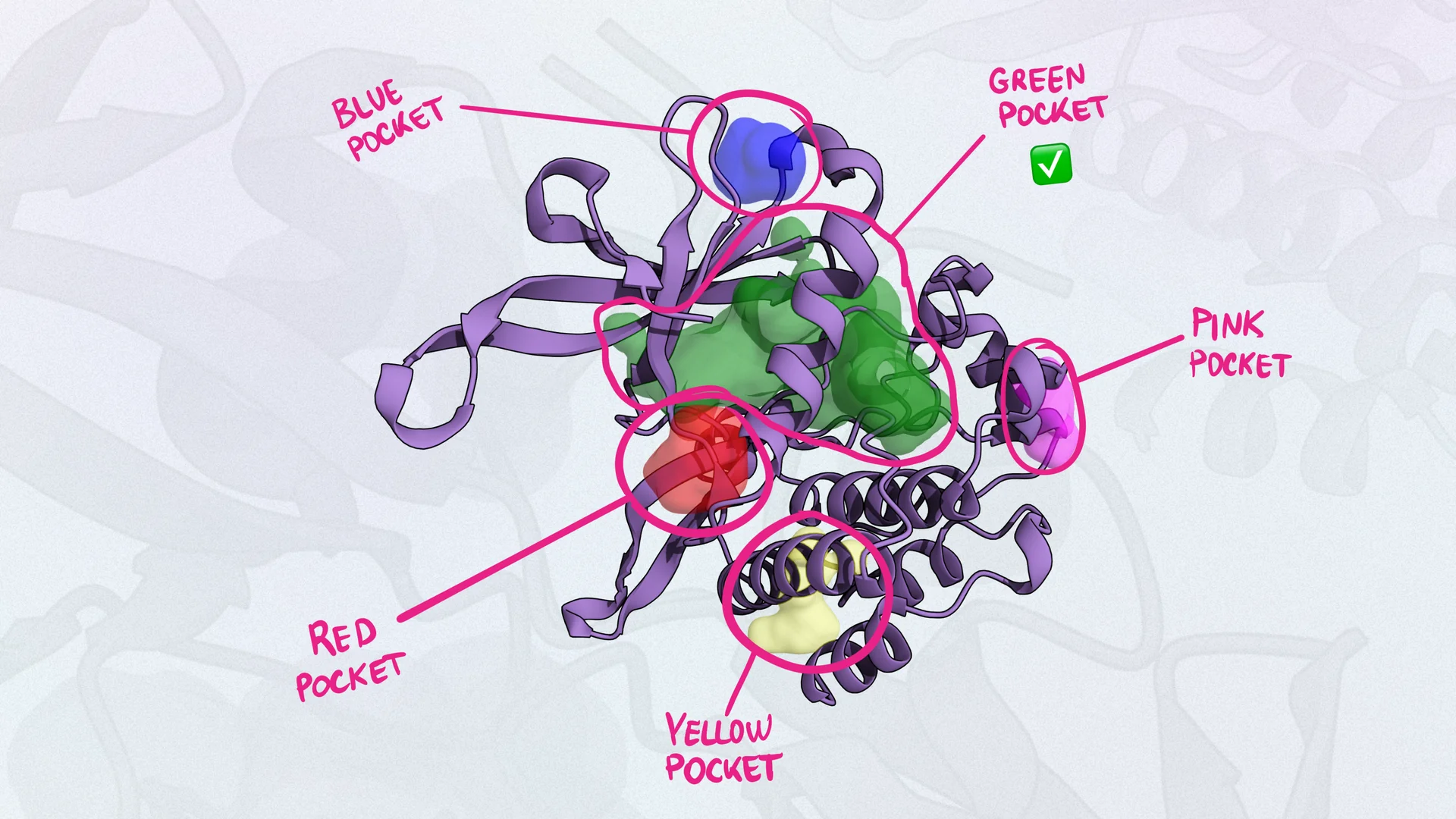
Ligand
definitionLigands are molecules that bind to specific sites on a target protein. If the binding affinity is strong enough, it often leads to a biological effect. Ligands can be endogenous (originating from within the body) or exogenous small molecules, peptides, or even larger proteins.
Importance of ligands in Computational Drug Discovery
- Target Interaction: Ligands interact with proteins to modulate their function, making them essential for understanding biological pathways and disease mechanisms.
- Binding Affinity: The strength of the interaction between a ligand and its target is crucial for therapeutic efficacy. Computational tools can predict binding affinities, guiding drug design.
- Selectivity: Computational methods can help optimize ligands to enhance binding, selectivity and their bioavailability, which can ultimately lead to better therapeutic outcomes.
- Optimization: Ligands should selectively bind to the target(s) of interest while avoiding other proteins, minimizing side effects.
Key Tools
-
AutoDock: Molecular docking software for predicting ligand-target interactions and binding.
-
BindingDB: Database for measured binding affinities of drug-like molecules.
-
ChEMBL: Database of bioactive molecules and their binding affinities, functional readouts or bioavailability.
-
DeepOrigin’s Ligand prep and property tools: Tools for predicting binding affinities, docking poses, and various molecular properties. Available in Balto and via API.
Literature
- “Using photolabile ligands in drug discovery and development”
-
Publication Date: 2000-02-01
-
Summary: Discusses three approaches: photoaffinity labeling, photoactivation and release of ‘caged ligands’, and photoimmobilization of ligands onto surfaces.
- “Fragment databases from screened ligands for drug discovery (FDSL-DD)”
-
Publication Date: 2023-11-01
-
Summary: Introduces a new method (FDSL-DD) that incorporates fragment characteristics into the drug development process.
- “Virtual Screening Algorithms in Drug Discovery: A Review Focused on Machine and Deep Learning Methods”
-
Publication Date: 2023-05-05
-
Summary: Reviews algorithms for virtual screening, highlighting the use of machine and deep learning
- “The Opportunities and Challenges of Peroxisome Proliferator-Activated Receptors Ligands in Clinical Drug Discovery and Development”
-
Publication Date: 2018-07-27
-
Summary: Provides an analysis of 84 types of PPAR ligands and their applications in clinical drug discovery.
- “Sustainable Drug Discovery of Multi-Target-Directed Ligands for Alzheimer’s Disease”
-
Publication Date: 2021-04-08
-
Summary: Reports the development of sustainable MTDLs derived from cashew nutshell liquid for Alzheimer’s disease.
- “Molecular Docking: Principles, Advances, and its Applications in Drug Discovery”
-
Publication Date: 2022-09-22
-
Summary: Reviews the principles and advances in molecular docking and its applications in drug discovery.
- “NMR in drug discovery: A practical guide to identification and validation of ligands interacting with biological macromolecules”
-
Publication Date: 2016-11-01
-
Summary: Introduces the concept of the validation cross to categorize experiments based on information content.
- “Machine learning approaches and their applications in drug discovery and design”
-
Publication Date: 2022-04-15
-
Summary: Reviews machine learning approaches used in chemoinformatics for drug discovery.
- “Network pharmacology: the next paradigm in drug discovery”
-
Publication Date: 2008-11-01
-
Summary: Discusses the role of polypharmacology in tackling efficacy and toxicity in drug development.
- “Advances in covalent drug discovery”
-
Publication Date: 2022-08-25
-
Summary: Reviews KRAS(G12C) inhibitors and discusses ligand-first and electrophile-first strategies.
- “Spotting and designing promiscuous ligands for drug discovery”
-
Publication Date: 2016-01-05
-
Summary: Discusses a computational tool for identifying promiscuous drug-like ligands.
- “What is the current value of MM/PBSA and MM/GBSA methods in drug discovery?”
-
Publication Date: 2021-06-24
-
Summary: Reviews methods for evaluating ligand-receptor interactions and binding free energy.
- “Merging Ligand-Based and Structure-Based Methods in Drug Discovery: An Overview of Combined Virtual Screening Approaches”
-
Publication Date: 2020-10-01
-
Summary: Analyzes hybrid LB + SB computational schemes in virtual screening.
- “Electrophilic warheads in covalent drug discovery: an overview”
-
Publication Date: 2022-02-06
-
Summary: Reviews electrophilic warheads used for protein labeling in chemical biology and medicinal chemistry.
- “Pharmacophore Modeling in Drug Discovery: Methodology and Current Status”
-
Publication Date: 2021-06-29
-
Summary: Discusses the advancements in pharmacophore modeling integrated with other computational methods.