Pharmacophore Modeling
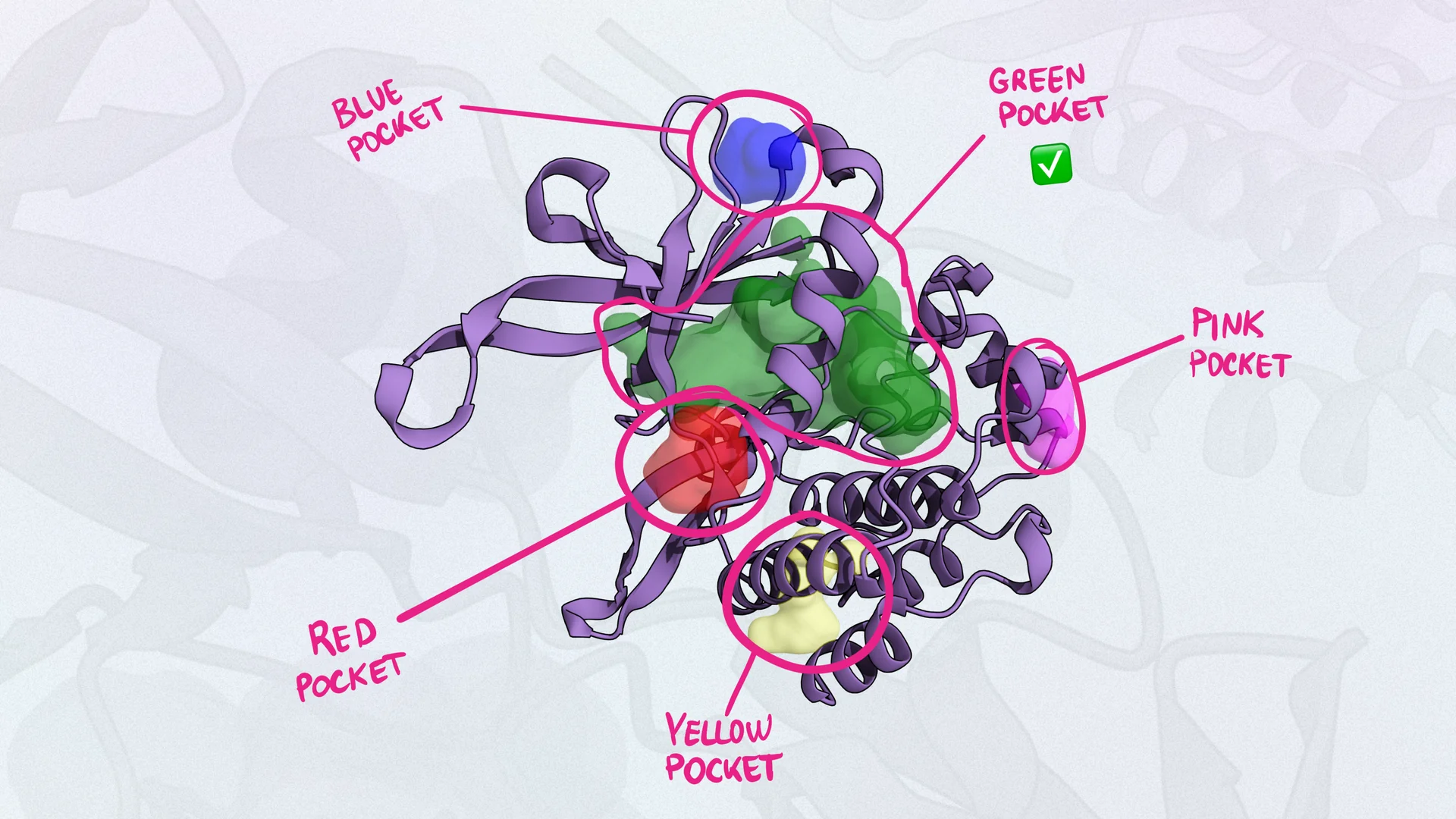
methodPharmacophore modeling is used to identify and represent the essential features of a molecule that are necessary for binding or activity. These features, known as pharmacophores, include hydrogen bond acceptors and donors, hydrophobic regions, aromatic rings, and charged groups. A pharmacophore model is a three-dimensional arrangement of these features that can be used to filter molecules to interact with a specific biological target to produce a desired effect.
Importance in Computational Drug Discovery
- Target Identification: Pharmacophore models help identify the key interaction points between a ligand and its biological target, facilitating the understanding of the molecular basis of ligand binding.
- Virtual Screening: Pharmacophore models can be used to screen large libraries of compounds to identify potential drug candidates that possess the required pharmacophoric features.
- Lead Optimization: By highlighting essential features for activity, pharmacophore models aid in the optimization of lead compounds to enhance their potency and selectivity.
Key Tools
-
Pharmit: An online platform for pharmacophore modeling and virtual screening.
-
LigandScout: A software tool for creating and applying pharmacophore models.
-
MOE (Molecular Operating Environment): A comprehensive suite for molecular modeling, including pharmacophore modeling and virtual screening.
-
Discovery Studio: A software platform offering tools for pharmacophore modeling, docking, and molecular dynamics simulations.
Literature
“Pharmacophore Modeling in Drug Discovery: Methodology and Current Status”
-
Publication Date: 2021-06-29
-
Summary: Reviews the methodology of pharmacophore modeling, its integration with other computational methods, and its applications in drug discovery.
“Pharmacophore Modeling in Drug Discovery and Development: An Overview”
-
Publication Date: 2007-02-28
-
Summary: Provides a historical overview of pharmacophore modeling and discusses developments in methodologies for pharmacophore identification and their applications in drug discovery.
“A Computer-Aided Drug Discovery Based Discovery of Lead-Like Compounds Against KDM5A for Cancers Using Pharmacophore Modeling and High-Throughput Virtual Screening”
-
Publication Date: 2021-10-12
-
Summary: Identifies lead compounds for KDM5A through pharmacophore modeling and high-throughput virtual screening, with further evaluation using ADMET properties and molecular dynamics simulations.
“The Development of Pharmacophore Modeling: Generation and Recent Applications in Drug Discovery”
-
Publication Date: 2018-12-08
-
Summary: Reviews successful examples of pharmacophore modeling applied in virtual screening and lead optimization, providing an overview of pharmacophore-based virtual screening.
“Pharmacophore Modeling: Advances, Limitations, and Current Utility in Drug Discovery”
-
Publication Date: 2014-11-11
-
Summary: Reviews the computational implementation of the pharmacophore concept and its common usage in drug discovery, including virtual screening, ADME-tox modeling, and target identification.
“Computational Discovery of SARS-CoV-2 NSP 16 Drug Candidates Based on Pharmacophore Modeling and Molecular Dynamics Simulation”
-
Publication Date: 2021-06-01
-
Summary: Uses pharmacophore-based virtual screening and molecular dynamics simulations to identify potential SARS-CoV-2 NSP 16 inhibitors.
“Azolium Analogues as CDK4 Inhibitors: Pharmacophore Modeling, 3D QSAR Study and New Lead Drug Discovery”
-
Publication Date: 2017-04-15
-
Summary: Presents ligand-based pharmacophore modeling and 3D-QSAR analyses for azolium-based CDK4 inhibitors.
“The Discovery of Novel BCR-ABL Tyrosine Kinase Inhibitors Using a Pharmacophore Modeling and Virtual Screening Approach”
-
Publication Date: 2021-03-04
-
Summary: Identifies novel BCR-ABL inhibitors through pharmacophore modeling and virtual screening, with in vitro validation.
“Unlocking Neuraminidase Inhibitors: Insights from Natural Products through Pharmacophore Modeling, Virtual Screening, and Molecular Docking”
-
Publication Date: 2024-11-08
-
Summary: Identifies potential neuraminidase inhibitors from natural products using pharmacophore modeling, virtual screening, and molecular docking.